CMN Weekly (10 July 2022) - Your Weekly CRISPR Medicine News
By: Karen O'Hanlon Cohrt - Jul. 10, 2022
Do you plan to organise a webinar?
Why not use the CMN platform and get exposed to the global gene editing community.
More details here.
Top picks
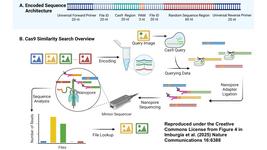
- In an article published in Nature Communications yesterday, scientists in China report the synergetic effect of CRISPR-activated graphene biointerfaces, and report the design of an on-line wearable microneedle patch for extraction and in vivo long-term monitoring of universal cell-free DNA (cfDNA). In their study, the team showed that the new system enables real-time monitoring of Epstein-Barr virus, sepsis, and kidney transplantation cfDNA with satisfactory sensitivity for 10 days in vivo. Data obtained from immunodeficient mouse models further demonstrated the feasibility and practicability of the proposed method, which the authors suggest holds great promise for long-term in vivo monitoring of cfDNA with potential for early disease screening and prognosis.
- In an article recently published in Science Advances, scientists in California and Brazil propose nickase-mediated homologous chromosome-templated repair (HTR) as a new and efficient and unanticipated mechanism for allelic correction. Specifically, the team showed that a Drosophila system performs efficient somatic repair of both double-stranded (DSBs) and single-stranded breaks (SSBs) using intact sequences from the homologous chromosome in a process they refer to as HTR. Unexpectedly, they found that HTR-mediated allelic conversion at the white locus was more efficient (40 to 65%) in response to SSBs induced by Cas9-derived nickases D10A or H840A than to DSBs induced by fully active Cas9 (20 to 30%). They also report that repair phenotypes elicited by nickase versus Cas9 differ in both developmental timing (late versus early stages, respectively) and the production of undesired mutagenic events (rare versus frequent). The authors propose that nickase-mediated HTR has far-reaching potential applications in the field of gene editing.
Research
- A team of scientists from various institutes in Boston recently showed that insertions of LINE-1 (L1) retrotransposons (RT) can occur frequently at CRISPR-Cas9 editing sites. Together with PolyA-seq and an improved amplicon sequencing, the team characterised more than 2,500 de novo L1 insertions at multiple CRISPR-Cas9 editing sites in human cell lines, and found that these L1 retrotransposition events exploit CRISPR-Cas9-induced double-stranded break formation and require L1 RT activity. They found de novo L1 insertions to be rare during genome editing by prime editors (PE), cytidine or adenine base editors (CBE or ABE), which is consistent with reduced DSB formation by these genome editors. The data demonstrate that RT insertions might be a potential outcome of CRISPR-Cas9 genome editing and provide further evidence on the safety of different CRISPR-based editing tools. The findings were published in Nature Communications.
- Until recently, very little was known about the potential of genome-editing technologies to help eradicate larger problematic mammals. To address this void, a team of researchers from the University of Adelaide in Australia developed a mathematical model capable of simulating the impact of gene drives on mammal populations at a landscape scale. Their findings, which were recently published in NeoBiota, represent the first estimates of times to eradication for long-lived alien mammals, and reveal that demography and life-history traits interact to determine the scalability of gene drives for vertebrate pest eradication. The authors note that optimism around eradicating smaller-bodied pests such as rodents and rabbits with gene-drive technologies does not easily translate into eradication of larger-bodied alien species such as cats and foxes.
- A team of researchers from multiple institutes in the U.S. report DNA Typewriter, a new system for in vivo molecular recording that overcomes the limitations of contemporary DNA-based memory devices. For DNA Typewriter, the blank recording medium (‘DNA Tape’) consists of a tandem array of partial CRISPR–Cas9 target sites, with all but the first site truncated at their 5′ ends and therefore inactive. Short insertional edits serve as symbols that record the identity of the prime-editing guide RNA mediating the edit while also shifting the position of the ‘type guide’ by one unit along the DNA Tape, that is, sequential genome editing. In their proof-of-concept study, the team demonstrate recording and decoding of thousands of symbols, complex event histories and short text messages. The full findings were published in Nature this week.
- Scienists in the UK and Sweden describe INDUCE-seq, which simultaneously detects the presence of low-level endogenous double-stranded breaks (DSBs) caused by physiological processes, and higher-level recurrent breaks induced by restriction enzymes or CRISPR-Cas nucleases. INDUCE-seq exploits an innovative next-generation sequencing (NGS) flow cell enrichment method, permitting the digital detection of breaks. It can therefore be used to determine the mechanism of DSB repair and to facilitate safe development of therapeutic genome editing. The authors also discuss how the method can be adapted to detect other genomic features. The findings were published yesterday in Nature Communications.
- In an article published in Molecular Therapy this week, a team in Denmark describes a truncated reverse transcriptase (RT) that enhances prime editing by split adeno-associated virus (AAV) vectors. Specifically, the team made advancements to the RT moiety of a prime editor (PE) to improve editing efficiencies and truncations to mitigate issues with AAV viral vector size limitations, which currently do not support efficient delivery of the large prime-editing components. Their efforts yielded a codon-optimised and size-minimised PE that has an expression advantage (1.4 × fold) and size advantage (621 bp shorter), which yields superior lentiviral vector titers (46 × fold) over the regular PE in an all-in-one PE lentiviral vector. Finally, delivery of the minimised PE to mouse liver by dual AAV8 vectors revealed up to 6% precise editing of the PCSK9 gene, thereby demonstrating the value of this truncated split PE system for in vivo applications.
- KRAS is the most frequently mutated oncogene in human cancers, and its activating mutations represent long-sought therapeutic targets. Researchers in Germany have now shown that cleavage of a panel of KRAS driver mutations suppresses growth in various human cancer cell lines, revealing their dependence on mutant KRAS, but that analysis of the remaining cells after long-term Cas9 expression revealed the occurence of oncogenic KRAS escape variants that were refractory to Cas9 cleavage. The team found that using an adenine base editor (ABE) to correct the oncogenic KRAS mutations progressively depleted the targeted cells without the appearance of the escape variants, allowing efficient and simultaneous correction of a cancer-associated TP53 mutation. The team demonstrated oncogenic KRAS and TP53 base editing in patient-derived cancer organoids. The findings, which were published in Cancer Research this week, suggests that base-editor-mediated correction of oncogenic mutations could be exploited in a personalised manner for future precision oncology applications.
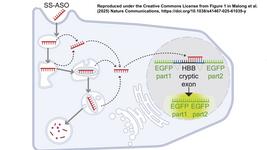
- Scientists in the UK and Sweden exploit the DNA cargo capacity of baculovirus to circumvent the delivery challenge in human cells. By encoding Cas9, sgRNA and donor DNAs on a single, rapidly assembled baculoviral vector, the team achieved whole-exon replacement in the intronic β-actin (ACTB) locus, including site-specific docking of very large DNA payloads, with up to 30% efficacy. They then used the approach to rescue wild-type podocin expression in steroid-resistant nephrotic syndrome (SRNS) patient-derived podocytes. The team demonstrated single baculovirus-vectored delivery of single and multiplexed prime-editing toolkits, achieving up to 100% cleavage-free DNA search-and-replace interventions without detectable indels. Their findings were published in Nucleic Acids Research earlier this week.
Reviews
Chemical Modifications of CRISPR RNAs to Improve Gene-Editing Activity and Specificity. The author of this review argues that the complexity of CRISPR presents rich unprecedented opportunities for chemists to synergise advances in synthetic methodology and structural biochemistry to rationally optimise crRNA-protein interactions.
Miscellaneous
- Open call for submissions to Frontiers in Genome Editing. Topic editors within therapeutic gene correction strategies are now welcoming the submission of articles that aid the basic understanding of precise genome editing and facilitate the clinical translation and therapeutic implementation of these tools in blood disorders or other monogenic diseases. Deadline 31st August 2022. See here for more details.
News from CRISPR Medicine News
- This week, we introduced our CMN disease overview, which will provide information on all the diseases that are targeted by gene-editing therapies involving CRISPR, prime editors, base editors, ZFNs, TALENs, Cas-CLOVER, MegaTAL and MegaNucleases. See the full list of diseases in our overview so far here.
- Don't miss our overview of the companies to follow within the gene-editing field. The list is growing all the time. Check out the overview here.
To get more of the CRISPR Medicine News delivered to your inbox, sign up to the free weekly CMN Newsletter here.
Tags
CLINICAL TRIALS
IND Enabling
Phase I
Phase II
Phase III
Recurrent or Progressive High-grade Glioma, (NCT06737146)
Sponsors:
Suzhou Maximum Bio-tech Co., Ltd.
Sponsors:
Suzhou Maximum Bio-tech Co., Ltd.
IND Enabling
Phase I
Phase II
Phase III
Advanced Peritoneal Malignancies or Abdominal Metastatic Solid Tumors, (NCT06912152)
Sponsors:
Zhejiang University
Sponsors:
Zhejiang University
IND Enabling
Phase I
Phase II
Phase III