CMN Weekly (2 September 2022) - Your Weekly CRISPR Medicine News
By: Gorm Palmgren - Sep. 2, 2022
Top picks
- British researchers have developed a method for near leak-free, inducible expression of a polycistronic array containing up to 24 gRNAs from two orthogonal CRISPR/Cas systems. The technique can increase CRISPRai multiplexing capacity and target gene flexibility. In yeast, they demonstrate the toolkit by targeting 11 genes in central metabolism in a single transformation, achieving a 45-fold increase in succinic acid, which could be precisely controlled in an inducible manner.
- CRISPR-mediated modulation of intracellular kinase signalling can improve the stemness and function of tumour-infiltrating lymphocytes (TIL) for adoptive cell therapy. Chinese researchers demonstrated this by using CRISPR to knockout AKT1 and/or AKT2 (protein kinase B) in TIL isolated from cervical and ovarian cancers.
Research
- In vivo studies in mice demonstrate that base editing after coronary artery disease and reperfusion can ameliorate oxidative injury to heart tissues and improve cardiac function. Base editing disrupted oxygen activation of the protein kinase CaMKII that otherwise can lead to fibrosis and remodelling of the extracellular matrix. In vitro studies in human cardiomyocytes showed markedly reduced arrhythmia in base-edited cells.
- Researchers in South Korea have used CRISPR-Cas9 to knockout three TTN-genes in human induced pluripotent stem cells (iPSC). TTN mutations are the common genetic cause for various types of cardiomyopathies (e.g., dilated cardiomyopathy, hypertrophic cardiomyopathy, restrictive cardiomyopathy, and arrhythmogenic right ventricular cardiomyopathy) and skeletal myopathies.
- In vivo studies in mice by Portuguese researchers suggest that CRISPR-Cas9 knockout of receptor-interacting protein kinase 3 (RIPK3) is promising in ameliorating non-alcoholic fatty liver disease (NAFLD) progression. Ripk3 deficiency rescued several biomarkers of NAFLD. This included impairment in mitochondrial biogenesis, bioenergetics and function in CDAA diet-fed mice and fat-loaded hepatocytes.
Industry
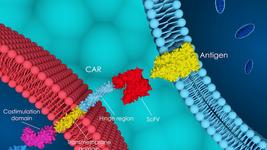
- ONK Therapeutics has presented promising in-vivo data of its optimized affinity CD38 CAR-NK candidate developed for treating multiple myeloma. The data showed potent anti-tumour activity for ONKT102 in-vitro and in-vivo in a CD38 positive MM.1S-LUC tumour model of multiple myeloma. ONKT102 is developed from expanded cord blood-derived NK cells that, among other modifications, have undergone CRISPR gene editing to knock out CD38 and prevent fratricide.
- The FDA has granted Intellia Therapeutics an orphan drug designation for its in vivo CRISPR-Cas9 genome editing candidate, NTLA-2002, for treating hereditary angioedema (HAE). NTLA-2002 is designed to knock out the target gene kallikrein B1 (KLKB1) to reduce plasma kallikrein activity and thus prevent HAE attacks.
Detection
- American researchers propose a system for enhanced CRISPR-Cas-mediated detection of nucleic acid and non-nucleic acid targets using enzyme-labelled reporters. The strategy offers a simple and sensitive platform to detect analytes where target amplification is either inconvenient (e.g., PCR under point-of-care settings) or impossible.
- A new detection system for Escherichia coli O157:H7 from beef samples is demonstrated by scientists from China. The system employs recombinase aided amplification (RAA) assisted CRISPR/Cas12a (RAA-CRISPR/Cas12a) fluorescence detection. It has a limit of detection of 5.4 × 102 CFU/mL, and detection was completed within 30 min after four hours of enrichment in spiked ground beef samples.
- Scientists in China have developed a highly sensitive and rapid platform for detecting SARS-CoV-2 via a portable CRISPR-Cas13a-based lateral flow assay. The limit of detection using CRISPR was 0.25 copy/μL, and results revealed 100% consistency.
- A highly sensitive detection system of cocaine employs a fluorescent aptasensor based on CRISPR-Cas12a combined with TdT. The detection limit is down to 33 pM, and the linear range is from 330 to 1.65 × 105 pM. Moreover, it can be successfully applied to the determination of cocaine in human plasma samples.
- Chinese researchers present a CRISPR-based method for amplified detection of therapeutic monoclonal antibodies. The technique uses target-triggered catalytic hairpin assembly activation of CRISPR/Cas12a and can detect panitumumab with a 0.62 pM detection limit.
- A Cas9 nickase variant with faster kinetics and enhanced targeting specificity is employed in an isothermal amplification reaction method using Cas9 nickase (Cas9 nAR) to detect genomic DNA. The technique was demonstrated by detecting several bacteria and viruses with a detection limit of fewer than ten copies of RNA molecules per reaction (20 μL volume).
Reviews
- Researchers in Thailand have looked at how mutations in different variants of Sars-CoV-2 (including Alpha, Beta, Gamma, Delta, and Omicron strains) could introduce mismatches to the previously reported primers and crRNAs used in the CRISPR-Cas system. They find that over 40% of the primer sets and 15% of the crRNAs contain mismatches.
- The potential of CRISPR-Cas9 prime editing for cardiovascular disease research and therapy is discussed in a review by American researchers. The study covers improvements in methods for in-vivo delivery of the prime editing components that should enable this technology to be used to edit the genome in patients.
News from CRISPR Medicine News
- In a Clinical Update on Wednesday, we reported that Graphite Bio had dosed its first patient with a potential CRISPR cure for sickle cell disease. The therapeutic strategy behind nula-cel is that CRISPR-based correction of the beta-globin (HBB) gene will decrease sickle haemoglobin (HbS) production and restore adult haemoglobin (HbA) expression. This can potentially cure SCD by restoring completely normally-functioning red blood cells.
To get more of the CRISPR Medicine News delivered to your inbox, sign up to the free weekly CMN Newsletter here.
Tags
CLINICAL TRIALS
IND Enabling
Phase I
Phase II
Phase III
IND Enabling
Phase I
Phase II
Phase III
Amyotrophic Lateral Sclerosis, ALS, or Frontotemporal Dementia FTD, (NCT04931862)
Sponsors:
Wave Life Sciences Ltd.
Sponsors:
Wave Life Sciences Ltd.
IND Enabling
Phase I
Phase II
Phase III