Clinical Trial Update – CRISPR-Based Diagnostics for Sars-CoV-2 and Mycobacterium
CMN Intelligence - The World’s Most Comprehensive Intelligence Platform for CRISPR-Genomic Medicine and Gene-Editing Clinical Development
Providing market intelligence, data infrastructure, analytics, and reporting services for the global gene-editing sector. Read more...
Multiple CRISPR-based therapies for blood diseases, cancer, diseases of the eye, and metabolic disorders are now in clinical development, and recently we have seen CRISPR being used as a diagnostic tool as well. A lot of the diagnostic efforts have centered around Covid-19 and here at CRISPR Medicine News we have looked into some of them in the past. Today, we present details from the clinical trials of two of these Sars-CoV-2 diagnostic tests, namely SHERLOCK by Sherlock Biosciences and DETECTR by Mammoth Biosciences.
Moreover, we present details from the clinical trial of a CRISPR-based diagnostic test for Mycobacterium tuberculosis that causes pulmonary tuberculosis and is developed at the Huashan Hospital in China. It is somehow an interesting co-incidence that the diagnostic tests concern these two diseases. In May 2020, a paper from an Indian research group suggested that Sars-CoV-2 can activate dormant tuberculosis infections through a temporary immunosuppressive effect.
SHERLOCK Diagnostic Kit for Sars-CoV-2
Sars-CoV-2 is a positive-sense single-stranded RNA coronavirus that causes Covid-19 in humans. It primarily infects cells in the respiratory system through binding to the receptor angiotensin-converting enzyme 2 (ACE2) and most often leads to symptoms like fever, dry cough, and fatigue. About 20% of those who get Covid-19, however, become seriously ill and require oxygen, with 5% becoming critically ill and needing intensive care.
Covid-19 is most commonly diagnosed by PCR analysis of swabs from the nose or throat to amplify viral genetic material to detectable levels and thereby confirm an active infection. Moreover, viral proteins can be detected using antibodies in several tests that are generally cheaper and quicker but less accurate. As an alternative to PCR and antigen tests, Sherlock Biosciences has developed a CRISPR-based test called SHERLOCK.
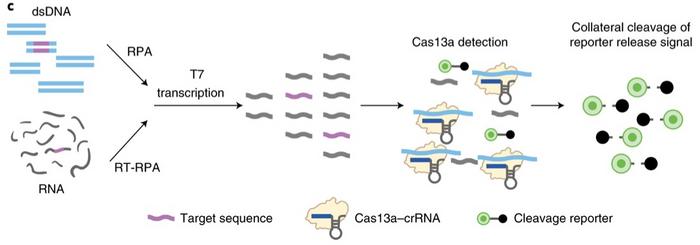
The SHERLOCK Diagnostic Kit for Sars-CoV-2 is based on collateral cleavage activity by CRISPR-Cas13. The method was described in detail in a Nature Biomedical Engineering paper in August 2020, and you can read our interview with Rahul Dhanda, CEO of Sherlock Biosciences here. The kit works by reverse transcribing viral RNA from respiratory swabs and then amplifying the DNA using the methods reverse transcriptase loop-mediated amplification (RT-LAMP) and strand-displacing DNA polymerase, respectively.
The amplified DNA is then transcribed and exposed to a CRISPR-Cas13 complex programmed to the target RNA sequences of the ORF1ab (O) gene and the Nucleoprotein (N) gene of SARS-CoV-2. When the sequences are found, non-specific activity results in cleavage of RNA reporters, resulting in a fluorescent readout detected by a plate reader. FDA granted the assay an Emergency Use Authorization (EUA) in May 2020.
You can read more about the SHERLOCK Diagnostic Kit for Sars-CoV-2 in our clinical trial database.
DETECTR Assay for Covid-19
The SARS-CoV-2 RNA DETECTR assay from Mammoth Biosciences is in some ways similar to SHERLOCK in that it uses RT-LAMP for amplification of viral genetic material and subsequent non-specific nuclease activity of a fluorescent marker upon target cleavage. However, DETECTR utilises CRISPR/Cas12 that targets single-stranded DNA. The method is described in detail in a Nature Biotechnology paper from April 2020.
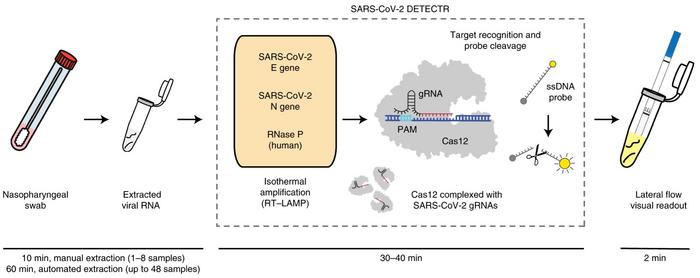
In DETECTR, CRISPR/Cas12 is programmed to target the Sars-CoV-2 envelope (E) and nucleoprotein (N) genes. It is a lateral flow assay that can detect SARS-CoV-2 from respiratory swab RNA extracts in less than 40 minutes. In July 2020, FDA granted the assay an Emergency Use Authorization (EUA).
You can read more about the DETECTR detection platform for Sars-CoV-2 in our clinical trial database.
CRISPR-based Mycobacterium tuberculosis Diagnostic Test for Pulmonary Tuberculosis
The bacterium Mycobacterium tuberculosis causes tuberculosis (TB). The bacterium usually attacks the lungs, but M. tuberculosis can attack any part of the body such as the kidney, spine, and brain. Not everyone infected with M. tuberculosis becomes sick. As a result, two TB-related conditions exist: latent TB infection (LTBI) and TB disease. If not treated properly, TB disease can be fatal. There is an estimated 10 million cases of TB worldwide annually leading to 1.5 million deaths every year. Both incidence and fatality have been slowly decreasing since 2000.
Two kinds of tests are used to detect TB bacteria in the body: the TB skin test (TST) and TB blood tests. A positive TB skin test or TB blood test only tells that a person has been infected with TB bacteria. It does not tell whether the person has latent TB infection (LTBI) or has progressed to TB disease. Other tests, such as a chest x-ray and a sample of sputum, are needed to see whether the person has TB disease.
The new CRISPR-based test for M. tuberculosis is based on the same principles ad the DETECTR Sars-CoV-2 test, i.e. it employs Cas12a detection of amplified M. tuberculosis sequences and subsequent collateral cleavage of a ssDNA fluorescent marker. Cas12a targets IS6110, which is an insertion element found exclusively within the members of the M. tuberculosis complex. The method is described in detail in an Emerging Microbes & Infections paper from September 2019.
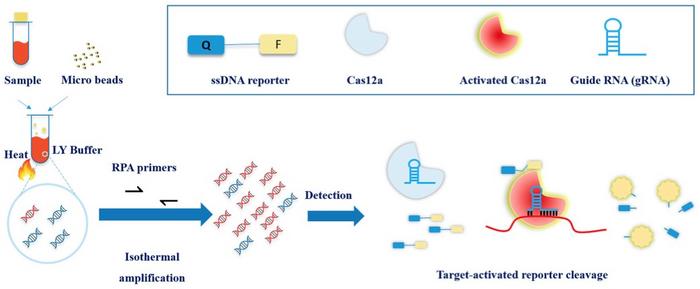
The tuberculosis diagnostic kit is now under clinical trial by Huashan Hospital in China, and they are currently recruiting 800 patients. It is an prospective observational study of pulmonary TB suspects who are undergoing sputum or bronchoalveolar lavage fluid (BALF) evaluation for pulmonary TB. The sensitivity and specificity of the CRISPR-based assay will be compared to clinical diagnosis, conventional culture methods and Xpert MTB/RIF assay on same batch specimens.
You can read more about the CRISPR-based Mycobacterium tuberculosis diagnostic test for pulmonary tuberculosis in our clinical trial database.
Tags
ArticleDiagnosticsCOVID-19Mycobacterium tuberculosisCRISPR-CasCas12aCas13aMammoth Biosciences, Inc.Sherlock Biosciences, Inc.TrialsApproved (FDA)
CLINICAL TRIALS
Sponsors:
Base Therapeutics (Shanghai) Co., Ltd.
Sponsors:
Base Therapeutics (Shanghai) Co., Ltd.