CMN Weekly (4 March 2022) - Your Weekly CRISPR Medicine News
By: Karen O'Hanlon Cohrt - Mar. 4, 2022
Top picks
- The years-long CRISPR patent dispute was settled this week, with the U.S. Patent and Trademark Office ruling that patents on the breakthrough gene-editing technology belong to Harvard University and the Massachusetts Institute of Technology, and not UC Berkeley.
- Gene editing biotechs face new uncertainty after CRISPR patent ruling. This piece in Biopharma Dive looks at what might happen for biotechs in the gene-editing space following the CRISPR patent verdict earlier this week.
- In an article published in Nature this week, researchers from The University of Texas at Austin describe a redesigned Cas9 that is 4,000 times less likely to make off-target edits without compromising targeted editing efficiency. The team used kinetics-guided cryo-electron microscopy to determine the structure of Cas9 at different stages of mismatch cleavage and used this data to specfically target regions of Cas9 that are exclusively involved in mismatch tolerance.
Research
- Researchers in Ireland demonstrate high levels of gene editing with 52.4% on-target insertions/deletions (indels) in primary hepatocytes following electroporation with CRISPR-Cas9 as mRNA and ribonucleoproteins (RNPs). The team also observed a transfection efficiency of >60% and on-target indels of up to 95% in primary mouse hepatocytes when targeting the Hpd gene, which encodes hydroxyphenylpyruvate dioxygenase. The findings, published in CRISPR Journal this week, present a potentially safe and efficient approach for therapeutic liver-directed gene editing and the production of liver disease models.
- Researchers in China generated and compared a set of C-to-G base editors (CGBEs) in human cells, and found that engineered deaminases rather than additional base excision repair proteins significantly improved C-to-G efficiency. They observed a significant increase in C-to-G transversions in the GC context when using rationally engineered eAID deaminase, and they could expand the targeting scope of CGBEs even further when using the SpRY Cas9 variant. The authors argue that these new CGBEs with engineered deaminase-nCas9 fusions broaden the base editing toolbox. The findings were published in CRISPR Journal this week.
- Scientists in the U.S. have developed a quantitative CRISPR sensing figure of merit (FOM) to compare different CRISPR-based nucleic acid sensor methods and to explore performance improvement strategies. The team define the CRISPR sensing FOM as the product of the limit of detection (LOD) and the associated CRISPR reaction time (T), and a smaller FOM means that the method in question can detect smaller target quantities faster. They then benchmarked the FOM performances of 57 existing studies, and among their findings they determined that digitalisation is the most promising amplification-free method for achieving comparable FOM performances to those using pre-amplification. The team argues that the findings would have broad implications for further optimisation of CRISPR-based sensing. The findings were published yesterday in ACS sensors.
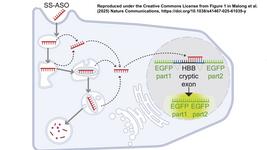
- Researchers in California report in Nature Communications this week that there is crosstalk between CRISPR-Cas9 and the human transcriptome. By performing eCLIP (enhanced cross-linking immunoprecipitation) of Cas9 in human cells, they found that Cas9 reproducibly interacts with hundreds of endogenous human RNA transcripts and that the association can be partially explained by a model built on gRNA secondary structure and sequence. They compare their findings to published genome-wide Cas9 DNA-binding or cut-site loci and published elevated transcriptome-wide RNA editing. The team reports that its findings do not support the hypothesis that human RNAs can broadly guide Cas9 to bind and cleave human genomic DNA, but that they illustrate a cellular and RNA impact likely inherent to CRISPR-Cas systems.
- Scientists in the U.S. present a newly-developed machine learning and computational tool, CASowary in BMC Genomics this week. They developed CASowary to predict the efficacy of a sgRNA using publicly available RNA knockdown data from Cas13 characterisation experiments for 555 sgRNAs targeting the transcriptome in HEK293 cells, in conjunction with transcriptome-wide protein occupancy information. According to their findings, CASowary can predict high quality guides for numerous transcripts in a cell line-specific manner.
- Researchers in California have systematically interrogated the ability of CRISPR-dCas9-based epigenome editors (Epi-dCas9) to engineer persistent epigenetic silencing. Their findings reveal that local chromatin features are important for the heritability of programmable silencing and the differential response to KRAB- and Ezh2-based epigenetic-editing platforms. The findings were published in Nucleic Acids Research this week.
Clinical news
- This week, Intellia Therapeutics and Regeneron announce long-term clinical data for their in vivotransthyretin (ATTR) amyloidosis candidate NTLA-2001. The data revealed deep and sustained reduction in disease protein levels following a single intravenous infusion of NTLA-2001. Read more in our clinical news update here.
- Intellia Therapeutics also announced this week that the first patient had been dosed in its Phase 1/2a clinical trial of NTLA-5001 for the treatment of acute myeloid leukaemia. Read more about NTLA-5001 in an earlier clinical update here.
Industry
- Editas Medicine announces favorable decision from U.S. Patent And Trademark Office In CRISPR patent interference. The patents at issue in the current interference are owned by the Broad Institute of Harvard and MIT and exclusively licensed to Editas Medicine for the development of therapies to treat serious diseases.
- Fate Therapeutics reports fourth quarter and full year 2021 financial results and highlights operational progress. The report includes a summary of progress on the company's iPSC-derived NK cell programmes for relapsed or refractory lymphoma as well as a status on its B cell cancer, acute myleloid leukaemia, multiple myeloma, and solid tumour programmes.
- This week, Cellectis provided a business update and reported fourth quarter and full year 2021 financial results. Highlights included a status on the ongoing BALLI-01 study for UCART22 for the treament of relapsed or refractory B-cell acute lymphoblastic leukaemia, as well as plans to submit an investigational new drug (IND) application for UCART20x22, the company's first allogeneic dual CAR T-cell product candidate for the treatment of B-cell non-Hodgkin’s Lymphoma.
Reviews
- Modeling genetic diseases in nonhuman primates through embryonic and germline modification: Considerations and challenges. This review in Science Translational Medicine discusses current methods and their limitations for producing reliable non-human primate genetic models of human disease. The authors reflect on how to ethically maximise the translatability of such models in the search for new therapeutic approaches to treating human diseases.
- Increasing the Targeting Scope of CRISPR Base Editing System Beyond NGG. In this review, authors in Canada discuss recent progress in the development of base editing, focusing on its targeting scope, and provide a workflow for selecting a suitable base editor based on the target nucleotide sequences.
News from CRISPR Medicine News
- On Monday we published 'PD-1 Targeted Cancer Immunotherapy Meets CRISPR'. For this interview, we spoke with Özcan Met PhD, who heads up the Cell Therapy Unit at Herlev and Gentofte Hospital in Denmark. Met and his colleagues recently integrated CRISPR gene editing into a clinical scale workflow for cancer immunotherapy. By knocking out the PD-1 protein with CRISPR-Cas9, they release a major brake on immune cells, and they believe that the approach could benefit certain cancer patients as early as next year. Read the interview here.
- In this week's clinical news update, we share news about Intellia Therapeutics' and Regeneron's ongoing Phase 1 trial for NTLA-2001 in transthyretin amyloidosis. Updated long-term clinical data demonstrates that the experimental in vivo CRISPR therapy resulted in rapid, deep and sustained responses in 15 patients. Read our news update here.
- The monthly CMN Markets Newsletter is here! CMN Markets February runs through Q4 2021 and full-year financial reports for Beam Therapeutics, CRISPR Therapeutics, Editas Medicine, Intellia Therapeutics and Allogene Therapeutics, and well as new deals and general market developments within gene-editing medicine. Read it here.
To get more of the CRISPR Medicine News delivered to your inbox, sign up to the free weekly CMN Newsletter here.
Tags
CLINICAL TRIALS
IND Enabling
Phase I
Phase II
Phase III
Recurrent or Progressive High-grade Glioma, (NCT06737146)
Sponsors:
Suzhou Maximum Bio-tech Co., Ltd.
Sponsors:
Suzhou Maximum Bio-tech Co., Ltd.
IND Enabling
Phase I
Phase II
Phase III
Advanced Peritoneal Malignancies or Abdominal Metastatic Solid Tumors, (NCT06912152)
Sponsors:
Zhejiang University
Sponsors:
Zhejiang University
IND Enabling
Phase I
Phase II
Phase III