CMN Weekly (21 April 2023) - Your Weekly CRISPR Medicine News
By: Karen O'Hanlon Cohrt - Apr. 21, 2023
CRISPR Medicine Conference 2024
- Have you seen that preparations are underway for the inaugural CRISPR Medicine Conference, to be held in Copenhagen in April 2024?? Check out the list of confirmed speakers here and don't forget to sign up for future updates by email.
Top picks
- Researchers in the U.S. show that prime editing can correct the sickle cell disease (SCD) allele (HBBS) to wild-type (HBBA) at frequencies of 15%-41% in haematopoietic stem and progenitor cells (HSPCs) from patients with SCD. Seventeen weeks after transplantation into immunodeficient mice, prime-edited SCD HSPCs maintained HBBA levels and displayed engraftment frequencies, haematopoietic differentiation and lineage maturation similar to those of unedited HSPCs from healthy donors. The findings were published earlier this week in Nature Biomedical Engineering.
- Microbiome-friendly phages join the campaign for better antimicrobials. This writeup in Nature provides an overview of some of the leading biotech companies that are exploiting natural and gene-edited bacteriophages to selectively combat drug-resistant bacterial infections without disturbing the host microbiome.
Research
- In an article published earlier this week in Nucleic Acids Research, a team of researchers in Germany describe a novel platform that consists of a Cas9 nuclease fused to DNA repair factors to synergistically inhibit non-homologous end joining (NHEJ) and favour homology-directed repair (HDR) for precise repairing of Cas-induced double-stranded breaks (DSBs). Compared to editing with canonical CRISPR-Cas9 methodology, the increase in error-free editing ranged from 1.5-fold to 7-fold in multiple cell lines and in primary human cells. The platform is compatible with a broad range of clinically-relevant repair templates and given its demonstrated safety benefit, the authors highlight its attractiveness as a tool for therapeutic applications that depend on precision genome editing.
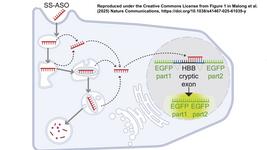
- In an article published earlier this week in Nature Communications, a team in the UK and Sri Lanka describes a base-editing strategy that corrects the HbE mutation (which causes haemoglobin E (HbE) β-thalassaemia) either to wild-type or a normal variant haemoglobin (E26G) known as Hb Aubenas, both of which recreate the asymptomatic trait phenotype. They report editing efficiencies in excess of 90% in primary human CD34 + cells, and show editing of long-term repopulating haematopoietic stem cells (LT-HSCs) using serial xenotransplantation in immunodeficient mice. The team also explored methods to predict and detect off-target effects.
- Glutaric aciduria type I (GA-1) is an inborn error of metabolism with a severe neurological phenotype caused by the deficiency of glutaryl–coenzyme A dehydrogenase (GCDH), the last enzyme of lysine catabolism. Using a series of knockout experiments in mice, a group of researchers in the U.S. uncovered that toxic GA-1 catabolites in the brain originate from the liver, and that it was possible to rescue the brain and lethal phenotype of the GA-1 mouse model by two different liver-directed gene therapy approaches, one of which involved deletion of the aminoadipate-semialdehyde synthase (Aass) gene via adeno-associated virus (AAV)-mediated delivery of CRISPR-Cas9 and a single Aass-targeting gRNA. Aass deletion prevented flux through the lysine degradation pathway. The findings were published this week in Science Translational Medicine.
- A team in the U.S. has developed a universal computational protein design framework, termed UniDesign, for designing protein-nucleic acid interactions. As proof of concept, they applied UniDesign to decode the preferred protospacer adjacent motif (PAM) and PAM-interacting amino acids (PIAAs) interactions for eight Cas9 and two Cas12a proteins. Their findings reveal that given native PIAAs, the UniDesign-predicted PAMs are largely identical to the natural PAMs of all Cas proteins. Given natural PAMs, the computationally redesigned PIAA residues largely recapitulated the native PIAAs (74% and 86% in terms of identity and similarity, respectively). The findings were published earlier this week in Briefings in Bioinformatics.
- Taking advantage of SpCas9-NG, a modified Cas9 with broad PAM flexibility, a team of researchers in Japan set out to correct a disease mutation in a patient with severe haemophilia B. They first generated induced pluripotent stem cells (iPSCs) from a patient with haemophilia B (c.947T>C; I316T) and established HEK293 cells and knock-in mice expressing the patient's F9 cDNA. They then transduced the cytidine base editor (C>T), including the nickase version of Cas9 (wild-type SpCas9 or SpCas9-NG), into the HEK293 cells and knock-in mice through plasmid transfection and an adeno-associated virus vector, respectively. They found that base editing with SpCas9-NG but not wild-type SpCas9 successfully converted C to T at the mutation site in the iPSCs, with indications of phenotype rescue in vitro and gene correction in vivo in experimental mice. The findings were published earlier this week in Communications Medicine.
- Scientists in Japan report CARTRIDGE (Cas-Responsive Translational Regulation Integratable into Diverse Gene control), a method to repurpose Cas proteins as translational modulators in mammalian cells. In an article published this week in Nature Communications, they demonstrate that a set of Cas proteins efficiently and orthogonally repress or activate the translation of designed mRNAs that contain a Cas-binding RNA motif in the 5'-UTR. By linking multiple Cas-mediated translational modulators, the team designed and built artificial circuits, including logic gates, cascades, and half-subtractor circuits.
- To address shortfalls in existing off-target prediction methods, which they claim do not fully exploit gRNA and DNA sequence pair information, two researchers in China have designed a novel coding scheme, which considers the key features of mismatch type, mismatch location and the gRNA-DNA sequence pair information. Furthermore, they developed a transformer-based anti-noise model called CrisprDNT to solve the noise problem that exists in off-target data. The researchers find, via experimental results of eight existing datasets, that the method that includes the anti-noise loss functions is superior to available state-of-the-art prediction methods. CrisprDNT is available at https://github.com/gzrgzx/CrisprDNT. The study findings were published this week in Briefings in Bioinformatics.
Industry
- Caribou Biosciences presented new pre-clinical data on CB-012, an allogeneic anti-CLL-1 CAR-T cell therapy, at the 2023 American Association for Cancer Research (AACR) Annual Meeting earlier this week. CB-012 is being developed as a novel treatment for adults with relapsed or refractory acute myeloid leukaemia (r/r AML). CB-012 is engineered with five edits using Caribou's proprietary Cas12a chRDNA technology, which armours the CAR-T cells for improved anti-tumour activity through checkpoint disruption and immune cloaking. The pre-clinical data, which can be view in a poster here, demonstrate that CB-012 exhibits enhanced anti-tumour activity against established AML xenografts. Learn more about Caribou's Cas12a chRDNA technology in our recent interview with CSO Dr. Steve Kanner here.
- Allogene Therapeutics presented interim Phase 1 data on its TALEN-edited candidate ALLO-316 in renal cell carcinoma (RCC) at the AACR Annual Meeting earlier this week. The ongoing Phase 1 TRAVERSE study is enroling patients with advanced or metastatic RCC who have progressed on standard therapies that include an immune checkpoint inhibitor and a VEGF-targeting therapy. Emerging data from this trial demonstrate the potential of an AlloCAR T to treat CD70-expressing RCC. In this trial, ALLO-316 showed early anti-tumour activity with deepening responses over time. Learn more about ALL0-316 in our previous clinical trial update here.
- Also at the recent AACR meeting, Jon Terrett, PhD and Head of Research at CRISPR Therapeutics, presented pre-clinical data on the company's next-generation CRISPR-Cas9-engineered allogeneic CAR-T cell candidates for treatment of liquid and solid tumours. Specifically, the presentation covered CTX112 and CTX131, which incorporate novel edits that increase potency and efficacy in the treatment of lymphoid and solid tumours. The full presentation can be viewed here.
Detection
- In an article published yesterday in Scientific Reports, a team of scientists in various institutions in the U.S. report a new predictive algorithm for the design of degenerate Cas13a CRISPR RNAS (crRNAs) using lassa virus as a model of highly variable RNA target. The work addresses the systematic design of minimum crRNA sets for the detection of diverse RNA targets using sequence degeneracy, and a single degenerate crRNA set adhering to rules identified in the study could detect representatives of all Lassa lineages.
- A team in China proposes a monkeypox virus (MPXV) detection assay combining recombinase-aided amplification (RAA) and CRISPR-Cas12a. This assay targeted the highly conserved MPXV F3L gene and demonstrates a low detection limit (LOD) of 101 copies per μL. By leveraging the high specificity nature of RAA and CRISPR-Cas12a, the team rationally optimised probes and conditions to achieve high selectivity that differentiates MPXV from other orthopox viruses and current high-profile viruses. To facilitate on-site screening of potential MPXV carriers, the team also developed a kit integrating lateral flow strips, which permitted naked-eye MPXV detection with a limit of detection of 104 copies per μL. The findings were published this week in Analytical Methods.
- In an article published yesterday in Analytical Chemistry, scientists in China report a sensitive CRISPR-Cas12a-assisted immunoassay for small molecule detection in homogeneous solution. An active DNA (acDNA) modified with a specific small molecule serves as a competitor for antibody binding and an activator of CRISPR-Cas12a. Large-sized antibody binding with the acDNA probe inactivates the collateral cleavage activity of CRISPR-Cas12a due to a steric effect. In the presence of a free small molecule targets, the small molecule-modified acDNA is displaced from the antibody, triggering catalytic cleavage of DNA reporters by CRISPR-Cas12a, and strong fluorescence is generated. The assay was capable of detecting test small molecules down to picomolar levels, using streptavidin or antibody as recognition elements.
- A team in South Korea has developed a novel Cas12a-based multiplex detection system by designing blocker DNA complementary to reporter DNA, which enables the simultaneous detection of two genes with a single Cas protein in a single reaction. To prove the concept, they chose high-risk human papillomavirus (HPV) 16 and 18 as model targets and incorporated recombinase polymerase amplification (RPA) and transcription reactions to achieve high accuracy and sensitivity. The team validated the performance of the novel workflow in detecting genomic DNA from various cell lines and clinical samples from cervical cancer patients with high specificity. The findings were published in Biosensors and Bioelectronics.
Reviews
- Using traditional machine learning and deep learning methods for on- and off-target prediction in CRISPR/Cas9: a review. This review provides an overview and comparative analysis of traditional machine learning and deep learning models used in CRISPR-Cas9 workflows, as well as challenges and possible future investigations in the field of on- and off-target prediction. The authors highlight key research challenges and directions associated with target activity prediction, as well as recent advances in the sgRNA-DNA sequence encoding used in state-of-the-art on- and off-target prediction models.
- Viral Vectors, Exosomes, and Vexosomes: Potential Armamentarium for Delivering CRISPR/Cas to Cancer Cells. This article reviews the commonly-used viral- (such as lentivirus, adenovirus, and adeno-associated viruses), and non-viral vectors including exosomes, and in particular tumour-derived exosomes (TDEs), that have proven useful in the delivery of CRISPR-Cas editing reagents. The authors also present information on the combined use of viral vectors and exosomes, called vexosomes, which may represent one solution to overcoming the limitations of both delivery systems.
Funding
- Earlier this week at the TED Conference in Vancouver, a new research initiative focused on untapping the potential of the microbiome, which is to be led by Jennifer Doudna and Jill Banfield at the Innovative Genomics Institute, was announced. The initiative, “Engineering the Microbiome with CRISPR to Improve our Climate and Health,” will receive $70 million in funding from donors, making it the largest scientific project funded through The Audacious Project (an initiative housed at TED) to date. Read full details here.
Podcasts
- Big Picture Science Radio Show: CRISPR Mosquitoes. This podcast series produced at the SETI Institute engages the public with modern science research through smart and humorous storytelling. The latest episiode explores the possible consequences of CRISPR technology and can be heard here.
News from CRISPR Medicine News
- Recent studies, including one led by Drs. Christiano Alves & Ben Kleinstiver at Massachusetts General Hospital were the first to demonstrate the curative potential of base editing for the devastating neuromuscular disease SMA, which is caused by mutations in the SMN1 gene. By substituting a single nucleotide in the SMN2 gene, a paralog of SMN1, the teams observed precise base editing and increased SMN expression in cells derived from SMA patients and in vivo in a mouse model of SMA. Read more in our lastest interview with Drs Christiano Alves & Ben Kleinstiver here.
To get more of the CRISPR Medicine News delivered to your inbox, sign up to the free weekly CMN Newsletter here.
Tags
CLINICAL TRIALS
IND Enabling
Phase I
Phase II
Phase III
Recurrent or Progressive High-grade Glioma, (NCT06737146)
Sponsors:
Suzhou Maximum Bio-tech Co., Ltd.
Sponsors:
Suzhou Maximum Bio-tech Co., Ltd.
IND Enabling
Phase I
Phase II
Phase III
Advanced Peritoneal Malignancies or Abdominal Metastatic Solid Tumors, (NCT06912152)
Sponsors:
Zhejiang University
Sponsors:
Zhejiang University
IND Enabling
Phase I
Phase II
Phase III