Fast CRISPR 'lab-on-a-chip' detects COVID19 in just over half an hour

A novel approach to the detection of the SARS-CoV-2 virus using CRISPR and electrical field-driven microfluidics reduces some of the difficulties with other diagnostics. It can move from a raw sample to a result in only 35 minutes, according to a new paper in PNAS.
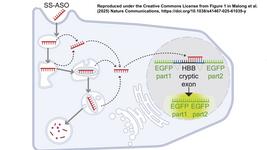
The use of CRISPR-based diagnostic approaches has garnered significant interest as the COVID-19 pandemic has spread across the world. The basic approach involves a CRISPR-Cas12 enzyme complexed with a synthetic guide RNA; that complex is activated when it binds to specific target DNA and cleaves it. It then non-specifically cleaves DNA reporter probes that are fluorescently labelled, producing a detectable signal.
»Our microfluidic approach to CRISPR-based diagnostics is distinct in that we perform key steps of sample preparation and the CRISPR detection process itself using electric fields - with no moving parts - applied within microchannels,« said Ashwin Ramachandran, of Stanford University in California, who led the new study.
Fast preparation and detection
Previous versions of a SARS-CoV-2 diagnostic, such as one published in Nature earlier this year, required more traditional sample preparation that could take as long as an hour.
There are several other diagnostics using CRISPR to detect COVID19. Researchers at Harvard University developed a fast system called CARMEN also using microfluidics and CRISPR. And we interviewed CEO ofSherlock BiosciencesRahul Dhanda about the world’s first FDA-authorized test.
The new approach involves the use of a selective ionic focusing technique called isotachophoresis (ITP) to perform automated purification of the target RNA from raw nasopharyngeal swabs. Next, an amplification step was done outside the microfluidic chip, but Ramachandran and the senior author, Juan G. Santiago, also of Stanford, said that they are working to incorporate that step into the chip as well.
The result of the amplification is reapplied to the microfluidics chip for the CRISPR-based part of the process. Electric fields, again using ITP, are applied in order to concentrate the Cas12-gRNA complex, the DNA reporter probes, and the target into a very small space - approximately 100 picoliters in volume, or around one-two-hundredth the size of a single grain of table salt.
»This dramatically accelerates the enzyme reactions and also focuses the fluorescence reporter molecules produced by one of these enzyme reactions,« Ramachandran said.
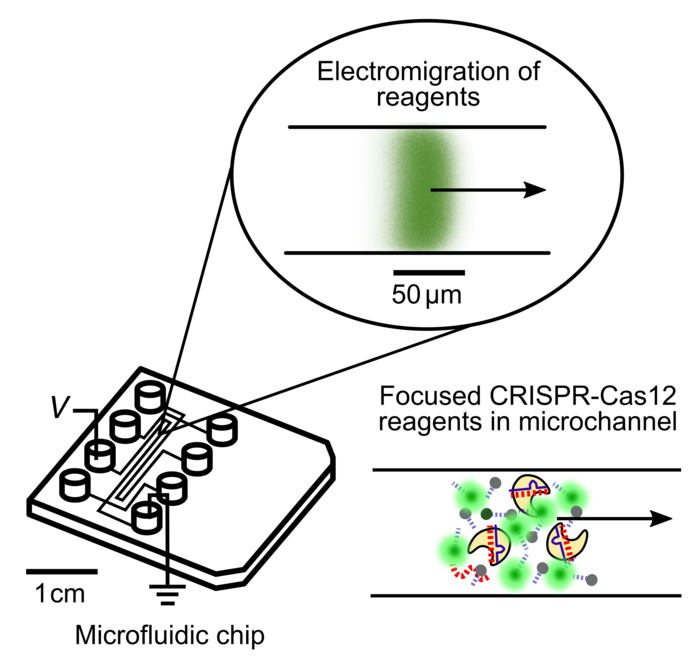
Colocating all the reagents into a tiny volume decreases the total amount of reagents needed by several hundred times. Also, it allows the CRISPR detection portion of the assay to finish in less than five minutes, compared to about 30 minutes for other CRISPR-based diagnostics. The fluorescent reporter molecules are also focused into a smaller area, which increases the strength of the detected signal.
Ramachandran and Santiago also pointed out that the process is highly automated and avoids both bulky equipment and manual steps associated with nucleic acid extraction, all of which could make the new approach attractive in the field.
Highly accurate with $5 price-tag
The group tested their process on a collection of 64 samples that had been tested using PCR.
The CRISPR-based approach successfully detected 30 of the 32 samples that were positive for SARS-CoV-2, and there were no false positives among the 32 negative samples, yielding positive and negative predictive agreements of 93.8% and 100%, respectively.
Juan G. Santiago said that the use of ITP allows them to focus the reagent ionic molecules of interest, avoiding other molecules. Specifically, they were able to increase the local concentration of the reactant molecules by about 1,000-fold.
»If you increase the concentration of reactants, you provide for the opportunity for a much higher rate of collisions between the molecules,« Santiago said.
»More collisions imply more opportunities for and higher rates of reaction.«
Fully integrated on a chip?
Others involved with CRISPR-based diagnostics weren't so sure it would be ready for prime time just yet.
»This method sounds great on paper, but the devil is in the details,« said Cameron Myhrvold, of the Broad Institute in the US, citing the need to perform some of the steps off the chip. »That is a lot of manipulation, and likely very challenging to automate due to the use of microfluidic chips,« Myhrvold said. »The approach is cool, and may have potential someday, but there is quite a bit of work to get from here to there.«
The group is working to fully integrate the steps of the process into a single microfluidic chip, avoiding the need to pipette off small volumes of extracted material for amplification before returning it to the chip. Once this has been achieved, they think the approach will be a very cheap method of SARS-CoV-2 - or other pathogens of interest - detection in the field.
»We anticipate that the cost of consumables (chip and reagents) will be less than $5 per sample,« Ramachandran said.
Dave Levitan is a science journalist based in the U.K.
Tags
ArticleInterviewNewsAcoustofluidicsDiagnosticsCOVID-19CRISPR-Cas
CLINICAL TRIALS
Sponsors:
Suzhou Maximum Bio-tech Co., Ltd.
Sponsors:
Zhejiang University