Adenoviral Vector Construction Gets Easier
Viral vectors are commonly used to deliver CRISPR components to human or other types of mammalian cells. Adenoviral vectors have several advantages over vectors based on, e.g., lentiviruses or adeno-associated viruses. These advantages include high carrying capacity and reduced off-target activity. The latter is caused by the fact that adenovirus neither integrates into the genome nor replicates during cell division, so Cas9 is only expressed transiently. However, the large adenovirus genome (~36 kb) and its limited number of restriction sites complicate the construction of recombinant adenoviruses by direct enzymatic ligation.
Several cloning strategies have been introduced to circumvent these challenges, and among them are homologous recombination in mammalian or bacterial cells. This approach avoids working with the sizeable adenoviral genome in vitro. Nevertheless, it involves time-consuming and laborious screening for homologous recombination and construction of multiple plasmids. This is a particular problem when working with the CRISPR-Cas9 system because several sgRNAs must usually be tested, thus requiring the construction of many different plasmids.
A recent paper in The CRISPR Journal by Gwangju Ji and co-workers from the Chinese Academy of Sciences in Beijing, China, presents a fast and simple system for adenoviral vector construction specially designed for CRISPR-Cas9 mediated gene editing. The system is called AdBlue, and it employs enzymatic target gene integration into the adenovirus genome without relying on specific restriction enzyme sites. As a result, the modified adenovirus genome is generated in just one hour in a one-step reaction. Furthermore, the selection of positive clones is efficient and accurate owing to combined kanamycin resistance and blue-white selection. All in all, only three days are required for vector construction before CRISPR-Cas9 experiments can be initiated, whereas the traditional system, AdEasy, requires five days.
Large DNA fragments are joined enzymatically
The AdBlue system is based on a method described by Daniel Gibson from the J. Craig Venter Institute in 2009. The method allows for the enzymatic assembly of DNA molecules up to several hundred kilobases in a single isothermal step. This is achieved by the concerted action of a 5′ exonuclease, a DNA polymerase and a DNA ligase. Briefly, two large DNA fragments sharing terminal sequence overlaps are joined into a covalently sealed molecule in a one-step isothermal reaction.
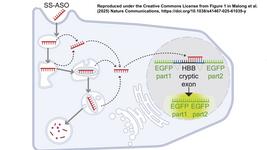
One of the fragments (pAdBlue) contains the adenoviral genome and the right inverted terminal repeat. The other fragment (pBlue) contains the gene of interest (Cas9 or sgRNA), selection genes and the left inverted terminal repeat. The terminal sequence overlaps are created by linearising the vectors with appropriate restriction enzymes, and subsequently, nucleotides are removed from the 5' ends by T5 exonuclease. Complementary single-stranded DNA overhangs are then annealed by DNA polymerase filling the gaps and DNA ligase sealing the nicks.
The resulting circular plasmid is transformed into TOP10 competent cells with over 90% transformation efficiency, and kanamycin-resistant transformants (blue spots) are selected and further amplified. The plasmid is then recovered and linearised, leaving a fragment encoding the adenoviral genome and the gene of interest surrounded by the left and right inverted repeat at either end. This fragment was then transfected into human embryonic kidney (AD-293) cells for virus packaging.
AdBlue is more efficient than AdEasy
Guang Ji and his colleagues first tested the efficiency of the vector construction system using adenoviruses that expressed enhanced green fluorescent protein (eGFP) as the gene of interest. A total of 24 blue colonies were selected on kanamycin-containing media, and genetic analyses revealed that 23 colonies (equivalent to 95.8%) contained the desired plasmid. These plasmids were then linearised and transfected into AD-293 cells for virus packaging. Expression of the gene of interest in host cells was shown by green fluorescence after 24 hours, and this was confirmed after five days by the appearance of green fluorescent plaques.
Compared to the traditional AdEasy system, the Chinese researchers found that AdBlue was more effective in obtaining recombinant adenoviruses. While AdEasy only selects positive clones by kanamycin resistance, AdBlue introduces blue-white screening and thus combines colour and kanamycin selection. This allows for much more efficient selection and reduces the number of clones to be analysed by a factor of ten, approximately. Moreover, the AdBlue system leads to relatively more positive plaques five days after transfection.
Finally, the research team tested the AdBlue system concerning CRISPR-Cas9. The approach is to deliver Cas9 and sgRNA in separate adenovirus particles, while other strategies combine the two in a single uniform virus particle. There are pros and cons to both systems. The separate system requires more time and labour, while it also has been shown to generate an up to 100 times higher functional virus titer than the uniform system.
CRISPR-mediated gene knockout is facilitated by AdBlue
Guangju Ji and co-workers evaluated gene knockout efficiency induced by adenoviral delivery of CRISPR-Cas9 nuclease complexes by targeting the sixth exon in the Celf1 gene. Two adenoviruses were produced expressing Cas9 and the sgRNA, respectively, and used for infection of cultured rat embryonic cardiomyocytes (H9C2) cells.
Seven days after infection, DNA disruption was observed downstream of the predicted cleavage site by Sanger sequencing, and insertion-deletion mutations (indels) were detected using the T7E1 nuclease assay. However, no DNA disruption or indels were detected in controls infected with Cas9 only and not sgRNA. Moreover, Western blot analysis revealed that Clef1 protein expression was significantly lower in cells transfected with both Cas9 and sgRNA compared to controls.
Based on these results, the Chinese researchers suggest that the novel AdBlue system can reduce the time and labour used to generate adenoviruses for the delivery of CRISPR-Cas9 complexes and induction of robust site-specific mutagenesis.
Link to the original article in The CRISPR Journal:
Tags
ArticleInterviewNewsDeliveryViralAdenovirus (AV)CRISPR-CasCas9
CLINICAL TRIALS
Sponsors:
Suzhou Maximum Bio-tech Co., Ltd.
Sponsors:
Zhejiang University