Engineered Virus-Like Particles Redefine CRISPR-Cas Delivery Options
CMN Intelligence - The World’s Most Comprehensive Intelligence Platform for CRISPR-Genomic Medicine and Gene-Editing Clinical Development
Providing market intelligence, data infrastructure, analytics, and reporting services for the global gene-editing sector. Read more...
Current in vivo genome-editing platforms each come with well-known constraints – limitations that eVLPs are designed to overcome. Adeno-associated viruses (AAVs) are clinically validated, but their packaging capacity of roughly 4.7 kb makes it difficult to deliver large editors such as base editors (BEs) or prime editors (PEs) in a single vector without resorting to split-vector strategies. Lipid nanoparticles (LNPs), now the most clinically advanced non-viral option, have made liver-directed CRISPR-Cas9 editing a clinical reality. However, they are still largely hepatotropic, and inefficient endosomal escape limits their usefulness in extrahepatic tissues.
eVLPs occupy a different structural niche. Rather than delivering DNA or mRNA, they package preassembled RNP complexes – the editor protein together with its guide RNA – and release them directly into the cytoplasm through virus-inspired membrane fusion. This approach bypasses endosomal trafficking and avoids sustained nuclease expression in recipient cells, resulting in a more transient editing window.
Structurally, eVLPs are nanoscale particles measuring 100–200 nm in diameter. They assemble around the retroviral Gag polyprotein, which forms an inner capsid core surrounded by a lipid envelope derived from the producer cell membrane. Production typically occurs in HEK293T cells via transient co-transfection with plasmids encoding Gag, a Gag–editor fusion, the guide RNA, and an envelope glycoproteinn (see Figure 1). Vesicular stomatitis virus glycoprotein (VSV-G) is commonly used to confer broad tropism, whereas alternatives, such as the baboon retroviral envelope (BaEV), can redirect targeting toward haematopoietic cells.
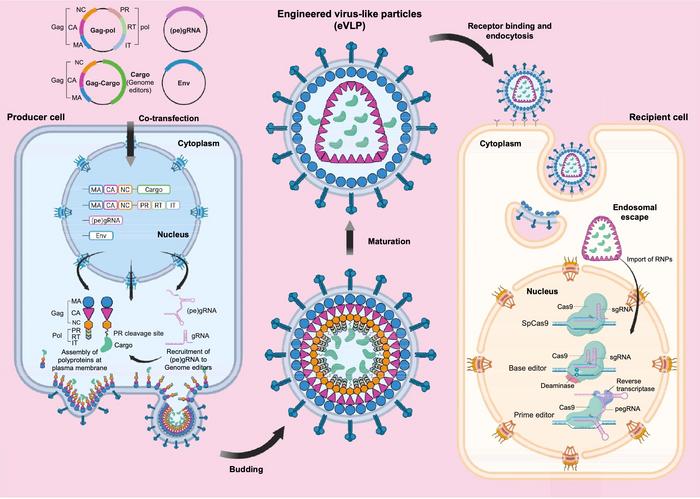
Throughout particle maturation, a co-packaged viral protease cleaves the linker between Gag and the cargo protein, releasing the RNP into the particle lumen. Upon contact with a recipient cell, the envelope mediates membrane fusion, the RNP is released into the cytoplasm, and intrinsic nuclear localisation signals guide it to the nucleus, where editing occurs.
The essential eVLP platform for CRISPR-Cas delivery – "Nanoblades", first described by Mangeot et al. in 2019 – used Moloney murine leukaemia virus (MMLV) Gag fused to Streptococcus pyogenes Cas9 (SpCas9). It demonstrated editing in primary cells and in mouse embryos in vivo. Subsequent development focused on systematically removing bottlenecks through successive BE-eVLP versions.
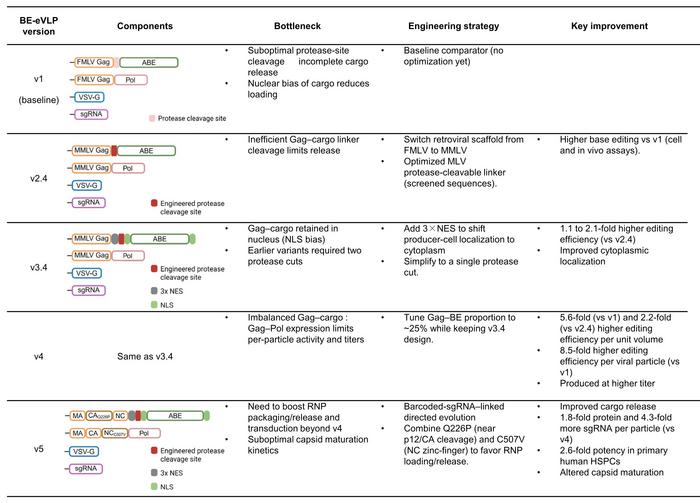
From v1 to v4, optimisation efforts included optimising the protease cleavage site to improve cargo release, adding a nuclear export signal (NES) to shift Gag–cargo fusions toward the cytoplasm during assembly, and adjusting the stoichiometric ratio of structural Gag to Gag–cargo plasmids to increase both the particle yield and RNP potency. By v4, a single intravenous injection achieved 63% editing of the Pcsk9 gene in mouse liver, resulting in a sustained 78% reduction in circulating PCSK9 protein levels. In the central nervous system, local administration produced up to 60% editing in transduced cells, while subretinal injection corrected the pathogenic Rpe65 mutation in the rd12 retinal degeneration model with efficiencies of up to 30% – sufficient to restore partial visual function.
The transition to v5 BE-eVLPs represented a shift from rational engineering to library-based directed evolution. Because eVLPs lack a viral genome, they do not naturally link genotype and phenotype as AAV capsid screens do. To address this, each Gag capsid mutant was tagged with a unique barcode embedded within the co-packaged sgRNA. Screening identified two cooperative substitutions – Q226P and C507V – that remodelled the capsid interior to better accommodate large RNP cargoes. Together, they delivered a two- to fourfold increase in editing potency over v4 and established a generalisable framework to enable further optimisation (see Figure 2).
Prime editing introduced additional complexity. The PE–reverse transcriptase fusion protein is substantially larger than base editors, and the prime editing guide RNA (pegRNA) contains a long 3′ extension that is particularly vulnerable to degradation. Simply adapting the v4 BE-eVLP design proved inadequate, resulting in the development of a dedicated PE-eVLP lineage.
In the v3b PE-eVLP, direct Gag–PE fusion was replaced with a smaller scaffold protein recruited via a coiled-coil interaction to improve cargo loading. The MS2/MCP aptamer system was substituted with the higher-affinity COM/Com pair to stabilise pegRNA capture. In vivo, a single subretinal injection corrected approximately 15% of the pathogenic Mfrp mutation in the rd6 model and around 7.2% of the Rpe65 substitution in rd12 mice, restoring partial visual function in both cases.
Parallel innovations have additionally expanded the platform. ENVLPE combined aptamer-mediated RNP recruitment, nucleocytosolic shuttling, and pegRNA stabilisation to achieve near-stoichiometric Cas9:pegRNA ratios and roughly 5.5-fold greater functional rescue in retinal models compared with earlier PE-eVLPs, at ten-fold lower particle doses. NanoScribes achieved near-transfection-level prime editing efficiencies and enabled editing in human induced pluripotent stem cells, hiPSC-derived haematopoietic stem cells, and primary myoblasts. The NanoMEDIC system delivered AsCas12a with Pol II-driven CRISPR RNA transcription, producing two- to threefold higher editing efficiencies than SpCas9-eVLPs and demonstrating CXCR4 knockout in T cells. The RIDE platform showed that tropism can be reprogrammed simply by exchanging the envelope plasmid – without altering the RNP payload – and demonstrated effective editing across ocular and neurological models, including initial tolerability data in non-human primates.
Pseudotyping flexibility is one of the platform’s core strengths. The DIRECTED approach separates fusogenic activity from receptor binding by disabling the native receptor-binding domain of VSV-G while retaining its membranefusion machinery. Cell specificity is conferred by interchangeable antibodies or ligands displayed on the particle surface, facilitating selective targeting of defined populations, such as human T and B cells. As the review notes, eVLPs function as "Trojan horse-like" delivery vehicles that combine virus-derived entry efficiency with the safety advantages of genome-free, transient cargo delivery, and are regarded as a complement to, rather than a replacement for, AAVs and LNPs.
Significant translational difficulties remain. GMP-compatible manufacturing has yet to be standardised, and current production relies on transient transfection of HEK293T cells, which introduces batch-to-batch variability. Systemic biodistribution data are limited, as most in vivo studies to date have relied on local administration pathways such as subretinal or intracerebral injection. Key questions also remain unresolved: the immunogenicity of retroviral structural proteins, the feasibility of repeat dosing, long-term genotoxicity risk, and the absence of non-human primate pharmacokinetic data. Till these gaps are addressed, a regulatory submission is unlikely.
The authors conclude that eVLPs are best viewed as a complementary platform. Their ability to deliver large and complex genome-editing RNPs in a transient, non-integrating manner makes them a notably compelling addition to the genome-editing delivery toolkit – provided that future efforts prioritise immune evasion, scalable bioprocessing, and rigorous safety evaluation alongside continued gains in editing efficiency.
The review was supervised by Hee Chan Yoo at the Chung-Ang University in South Korea. It was published in BioDrugs on 24 February 2026.
To get more CRISPR Medicine News delivered to your inbox, sign up to the free weekly CMN Newsletter here.
Tags
ArticleCMN HighlightsNewsDeliveryIn vivoVirus-like particle (VLP)
CLINICAL TRIALS
Sponsors:
Base Therapeutics (Shanghai) Co., Ltd.
Sponsors:
Base Therapeutics (Shanghai) Co., Ltd.