Explainer: DNA, RNA or Protein: Which Format Is Best for Your CRISPR Edit?
CMN Intelligence - The World’s Most Comprehensive Intelligence Platform for CRISPR-Genomic Medicine and Gene-Editing Clinical Development
Providing market intelligence, data infrastructure, analytics, and reporting services for the global gene-editing sector. Read more...
CRISPR has opened the doors to many possibilities within genome engineering and gene therapy. CRISPR-Cas gene editing technology relies on two critical components: a Cas endonuclease and a gene-targeting guide RNA (gRNA). The ultimate goal of a CRISPR experiment is to get a Cas enzyme and a target-specific gRNA to the nucleus of a cell to carry out the desired edit. Cas proteins and gRNAs may be transfected into cells in three formats, as outlined below and illustrated in Figure 1.
(1) DNA
- Two plasmids, one encoding Cas, the other encoding gRNA
- One plasmid encoding Cas and gRNA
- Dual viral vectors, one encoding Cas, the other encoding gRNA
- One viral vector encoding Cas and gRNA
(2) RNA
- Cas-encoding messenger RNA (mRNA) and gRNA
(3) Protein
- Purified Cas protein and gRNA
- Cas/gRNA ribonucleoprotein (RNP) complex
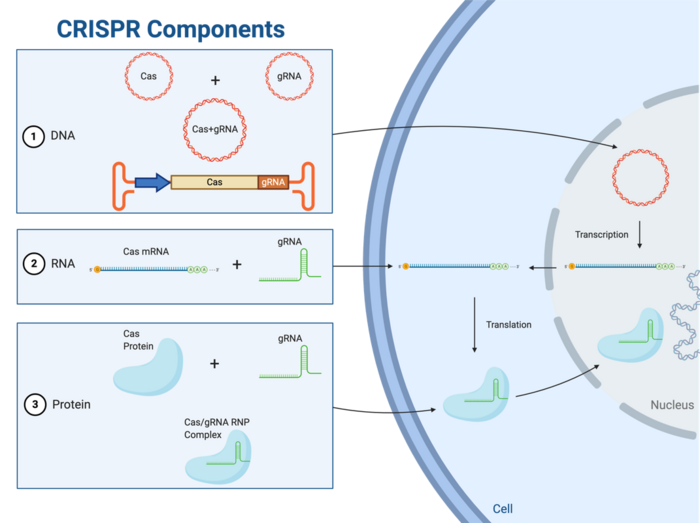
All Down to the Ribonucleoprotein (RNP) Complex
Regardless of which delivery format is used, the ultimate prerequisite for gene editing is the presence of a ribonucleoprotein complex, which can be thought of as the CRISPR gene-editing unit. Once present in the cell, the Cas endonuclease fuses with the gRNA in the cytoplasm to form the ribonucleoprotein (RNP) complex (Figure 1). The RNP complex must be able to access the nucleus for any edit to take place, and scientists have made this possible by engineering Cas-encoding genes to express a specific short sequence known as a nuclear localisation signal (NLS). The NLS signal is expressed as part of the Cas transcript and is translated to a tag on the Cas protein that is recognised by nuclear import proteins, thus facilitating Cas import into the nucleus.
The exact path that leads to formation of the RNP complex depends on the delivery format used to bring Cas and gRNA into the cells. This article will help you understand the basic advantages and limitations associated with each of these formats.
DNA Format
Plasmids
DNA is a highly stable molecule that is easy to produce in the lab at low cost and in large quantities. For these reasons, plasmids encoding Cas and the relevant gRNA may be an attractive choice for CRISPR. However, plasmids must be able to cross the plasma and nuclear membranes for expression and this can be challenging. Furthermore, plasmid-based delivery may lead to constitutive Cas and gRNA expression for extended periods of time, and while sustained expression might be advantageous in some gene editing scenarios, it can also overwhelm the cells with undesired off-target editing (1). Moreover, there is some evidence that plasmid delivery may trigger unwanted immune responses (2).
Viral Vectors
An alternative and preferred approach to DNA-mediated delivery is to engineer genes encoding the vital CRISPR components into a viral genome, and then transfect cells with this virus.
Within CRISPR and gene therapy, adeno-associated viruses (AAVs) are the most widely used viral vectors owing to their safety and transduction efficiency. One drawback with these vectors however is their limited packaging capacity of approximately 4.7 kbp (3). This creates a challenge when trying to package the especially large Cas gene and promoter (approximately 4.2 kbp, for the most commonly used Cas9 from Streptococcus pyogenes along with the gRNA. The packaging problem is made worse if a donor DNA template is to be co-introduced for homology-directed repair at the genomic cut site. To circumvent these packaging limits, researchers frequently use a dual AAV system, where one AAV encodes Cas9 and the other encodes the gRNA and donor DNA template (4). For successful CRISPR, both of these AAVs need to be in the same cell at the same time. Although not problematic for lab research, it is worth noting that viral capsids expressed by AAV and other viral delivery vectors may provoke an immune response in the host. This may be an obstacle in gene therapy because it poses the risk of patients becoming ‘immune’ to the therapy after repeated administration (5).
RNA Format
As an alternative to the DNA format, the CRISPR components can be delivered as Cas mRNA and gRNA. This approach circumvents the need to cross the nuclear membrane for transcription, since the Cas mRNA can be directly translated by ribosomes in the cytoplasm (Figure 1). However, RNA is a less stable molecule than DNA and is more susceptible to degradation from enzymes found in the laboratory as well as inside the cells/body. This has led to efforts within the CRISPR and gene therapy communities to develop chemically modified mRNAs or gRNAs with improved stability (6).
Using RNA as opposed to DNA delivery results in faster gene editing because there is no requirement for transcription. Another advantage of using this format is that RNA has a transient presence inside cells because of its inherent instability, which mitigates undesirable off-target editing (1). However, in a clinical setting the RNA format may also reduce the longevity of any therapeutic effect.
Ribonucleoprotein (RNP) Format
Functional, ready-to-use RNPhas become popular within the CRISPR community in recent years. Since the goal of all delivery formats is to ensure the presence of a RNP complex inside the nucleus of host cells, delivering the assembled complex directly to cells is highly attractive since it requires the least amount of intracellular processing (Figure 1).
Various methods exist for delivery RNPs to cells, with electroporation being the most common. Here, an electric field is applied to the cells in culture, which leads to the formation of small pores in the plasma membrane through which the RNP can pass. Once inside the cell, RNPs are quickly imported into the nucleus via NLS for genome editing. Thus, since RNP delivery bypasses the need for mRNA transcription and translation, gene editing happens relatively fast compared to other formats of delivery (7).
A RNP complex engulfs and protects the unstable gRNA from degradation, and its relatively short lifetime inside cells lowers the risk for off-target editing (6, 8). However, one must also consider that the degradation of RNPs over time may limit their suitability in scenarios that require long-term stable Cas expression. Additional limitations are the many steps involved in Cas expression and purification, which will require additional laboratory reagents, equipment, and expertise, and which is more costly than obtaining a CRISPR-encoding DNA or mRNA (1).
So, Which Format Is Best for Your CRISPR Edit?
The truth is that there is no one-size-fits-all answer, but we hope this overview will help you to decide which format is best for your needs. Although the DNA format is generally the easiest to handle, it may not always be optimal because of packaging limitations or risk of off-target editing. The RNA delivery format ensures a shorter experimental time, and transient expression of CRISPR components may be advantageous albeit with the caveat that inherent RNA instability may reduce overall editing efficiency. RNPs offer advantages such as speed and simplicity but are more costly and demanding to produce in the lab.
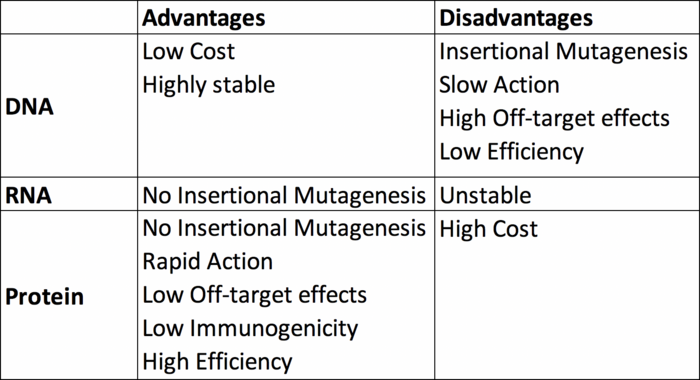
Finally, here’s a tabulated summary of the three delivery formats discussed here.
Karim Shalaby is a passionate science writer, preparing his PhD on CRISPR at the Biological and Biomedical Sciences Division, College of Health and Life Sciences, Hamad Bin Khalifa University, Doha, Qatar.
Literature Cited:
1. Z. Glass, M. Lee, Y. Li, Q. Xu, Engineering the Delivery System for CRISPR-Based Genome Editing. Trends Biotechnol36, 173-185 (2018).
2. T. S. Hughes et al., Immunogenicity of intrathecal plasmid gene delivery: cytokine release and effects on transgene expression. J Gene Med11, 782-790 (2009).
3. M. E. McClements, R. E. MacLaren, Adeno-associated Virus (AAV) Dual Vector Strategies for Gene Therapy Encoding Large Transgenes. Yale J Biol Med90, 611-623 (2017).
4. C. L. Xu, M. Z. C. Ruan, V. B. Mahajan, S. H. Tsang, Viral Delivery Systems for CRISPR. Viruses11, (2019).
5. H. C. Verdera, K. Kuranda, F. Mingozzi, AAV Vector Immunogenicity in Humans: A Long Journey to Successful Gene Transfer. Mol Ther28, 723-746 (2020).
6. A. Hendel et al., Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat Biotechnol33, 985-989 (2015).
7. A. K. Fajrial, Q. Q. He, N. I. Wirusanti, J. E. Slansky, X. Ding, A review of emerging physical transfection methods for CRISPR/Cas9-mediated gene editing. Theranostics10, 5532-5549 (2020).
8. S. Kim, D. Kim, S. W. Cho, J. Kim, J. S. Kim, Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res24, 1012-1019 (2014).
Tags
CLINICAL TRIALS
Sponsors:
Base Therapeutics (Shanghai) Co., Ltd.
Sponsors:
Base Therapeutics (Shanghai) Co., Ltd.







