Which Protein Domain is Best for CRISPRi? A New Study Has the Answer

What began as a PhD student’s side project could have important applications for future gene silencing therapies.
Nader Alerasool, a graduate student in Mikko Taipale’s lab at the University of Toronto, has identified a protein domain that, when fused to catalytically inactive Cas9 (dCas9), can strongly repress gene expression in human cells. The newly characterised repressor, called ZIM3, could prove useful in CRISPR screen experiments, in which dozens or thousands of genes are turned “off” to identify each gene’s role within a cell. Specifically, with ZIM3, researchers will be better equipped to find molecular targets to treat cancer, probe neurological diseases, and identify new drugs.
ZIM3 Is a Krüppel Associated Box (Krab) Domain
ZIM3 is one of hundreds of Krüppel associated box (KRAB) domains—transcriptional regulators found in zinc finger proteins—that are encoded in human cells. KRAB proteins repress gene expression because they recruit TRIM28/KAP1, a scaffold protein that binds with protein complexes involved in chromatin regulation.
The first study to use dCas9 for targeted gene regulation was published in 2013. Since then, most studies have used the KRAB domain from KOX1 for CRISPR inhibition (CRISPRi) experiments, likely because it was the first KRAB to be functionally characterised. In 2018, George Church’s group reported a rationally designed repressor domain, called KOX1 KRAB–MeCP2 which, when fused to dCas9, exhibited repression efficiencies far stronger than KOX1.
In his experiments, Alerasool fused 57 different KRAB domains to dCas9 and measured the repression efficiency for each one. It turned out that severalKRAB domains were better repressors than KOX1 and KRAB-MeCP2, and ZIM3 was the strongest repressor tested.
All of the KRAB domains analysed in Alerasool’s recent study, which was published earlier this month in Nature Methods, were tested in two human cell lines: embryonic kidney 293T (HEK293T) and K562 cells, an immortalised myelogenous leukaemic cell line that originates from bone marrow.
Both cell lines expressed green fluorescent protein (GFP) and were monoclonal, meaning that they are genetically identical. Alerasool used the same guide RNAs when testing each KRAB-dCas9 fusion protein to ensure consistent results across experiments.
We interviewed Mikko Taipale and Nader Alerasool to learn more about this work.
Initial Inspiration and Testing the KRABs
- Where did this idea come from?
Alerasool: Well, this idea wasn't really planned out. I was working on multiple projects and having some problems progressing with my PhD. Experiments weren't working as I had hoped, so I was working on many different things. The work in the new paper actually started as one of my side projects, because we collected interesting data where we saw that the KOX1 KRAB domain was not a very strong repressor amongst the ones that we tested. We thought, ‘Wow, that’s surprising!’ Then, in 2018, George Church’s group published the engineered KOX1-MeCP2 repressor, and we immediately wanted to benchmark their work to see if that was a stronger repressor.
Taipale: Tim Hughes’ lab also published a paper, on KRAB domains, zinc fingers, and their protein-protein interactions in 2016. We wanted to test how well KRAB domains would work as repressors. We were surprised that to find that KOX1 actually wasn't that strong as a repressor, but we figured that it's probably used for historical reasons, since that was the very first KRAB domain to be functionally described. And then people just continued with that one instead of looking for other KRAB domains. Then, when the Church paper came out, we were already testing KRAB domains with CRISPRi at that point. But in our hands, at least, ZIM3 seemed to work even better than the Church lab’s repressor or KOX1 alone.
CRISPR and dCas9 can also be used to activate genes. Read about how researchers are using this strategy in a attempt to cure heritary blindless without cutting DNA.
- Can you tell me a bit about the experiments?
Alerasool: We designed the reporter assay ourselves, but used reporter constructs from Stanley Qi’s group at Stanford from their 2016 Nature Methods paper.
First, I made clonal lines from the repressor reporter, which has a GFP-expressing construct that's randomly integrated into the genome. Each of the clonal lines also expressed a guide RNA. We then introduced 57 different KRAB-dCas9 genes into the cell lines [HEK293T and K562] by viral transfection, and measured the GFP levels using flow cytometry at different time points. ZIM3 was the strongest repressor.
What Makes ZIM3 So Effective?
- Do you know, mechanistically, why ZIM3 is such a strong repressor?
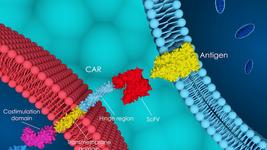
Alerasool: I can’t imagine ZIM3 being mechanistically different from the other KRABs. dCas9, when fused to KRAB domains, mediates repression by recruiting a co-repressor called TRIM28/KAP1. So we think that the mechanism is the same for all the KRAB domains we tested, but the repression strength is based on the affinity between the KRAB domain and TRIM28/KAP1.
Taipale: I mean, we have only correlations. So we can't know for sure that this is the case, but it's a very reasonable hypothesis.
- Was anything about this study surprising?
Taipale: This was a project that didn't have too many surprises. ZIM3 just seemed to work really well. It is surprising, though, that KOX1 didn't work that well. We had heard through the grapevine over the years that CRISPRi hasn't really worked that well in other people’s hands. And yet, you see all these publications that boast how well CRISPRi works. So I think there's definitely some publication bias there. If you just read the papers in this field, you might think that there’s not much room for improvement. But as we spoke with others, it was clear that a stronger repressor would be useful for CRISPRi applications.
- Each of the 57 different KRAB domains that you tested exhibited a slightly different repression efficiency. Do you think that having a range of characterised KRAB domains for CRISPRi is useful?
Taipale: It could be useful, yeah. At least in my view, for CRISPRi screens in human cells, it’s probably best to have something like ZIM3 that represses really well. But for other purposes, you might want to have something that tunes down gene expression just a little bit, or something that could tune down expression by 30 % or 40 %, instead of a complete shutdown. It would also be useful to have repression domains that can induce permanent heterochromatin versus temporary repression. That’s the way I see the field going forward with CRISPRi for therapeutic applications.
- What comes next for the lab?
Taipale: We’re planning to submit a manuscript soon on transcriptional activation, where we’ve used dCas9 to study transcriptional activators. We’re also following up on other KRAB domains and non-KRAB repressors as well.
Niko McCarty is a science journalist and former synthetic biologist. He lives in New York.
Link to the original article in Nature Methods:
An efficient KRAB domain for CRISPRi applications in human cells.
(Interview is condensed and edited for clarity).
FACT BOX: Taipale Lab
Mikko Taipale’s Lab is located in the Donnelly Centre at the University of Toronto. The lab’s research is organised around two main themes: development of large-scale genetics and ‘omics’ methods for biology, and studies that seek to understand how disruptions to protein homeostasis lead to human disease. The lab routinely uses techniques within molecular biology (including CRISPR assays), functional genomics and proteomics.
Tags
ArticleInterviewNewsDonnelly Centre for Cellular and Biomolecular Research, University of TorontoReagentsCas9dCas9CRISPRi
CLINICAL TRIALS
Sponsors:
Wave Life Sciences Ltd.